PAINS in the Assay: Chemical Mechanisms of Assay Interference and Promiscuous Enzymatic Inhibition Observed during a Sulfhydryl-Scavenging HTS | Journal of Medicinal Chemistry

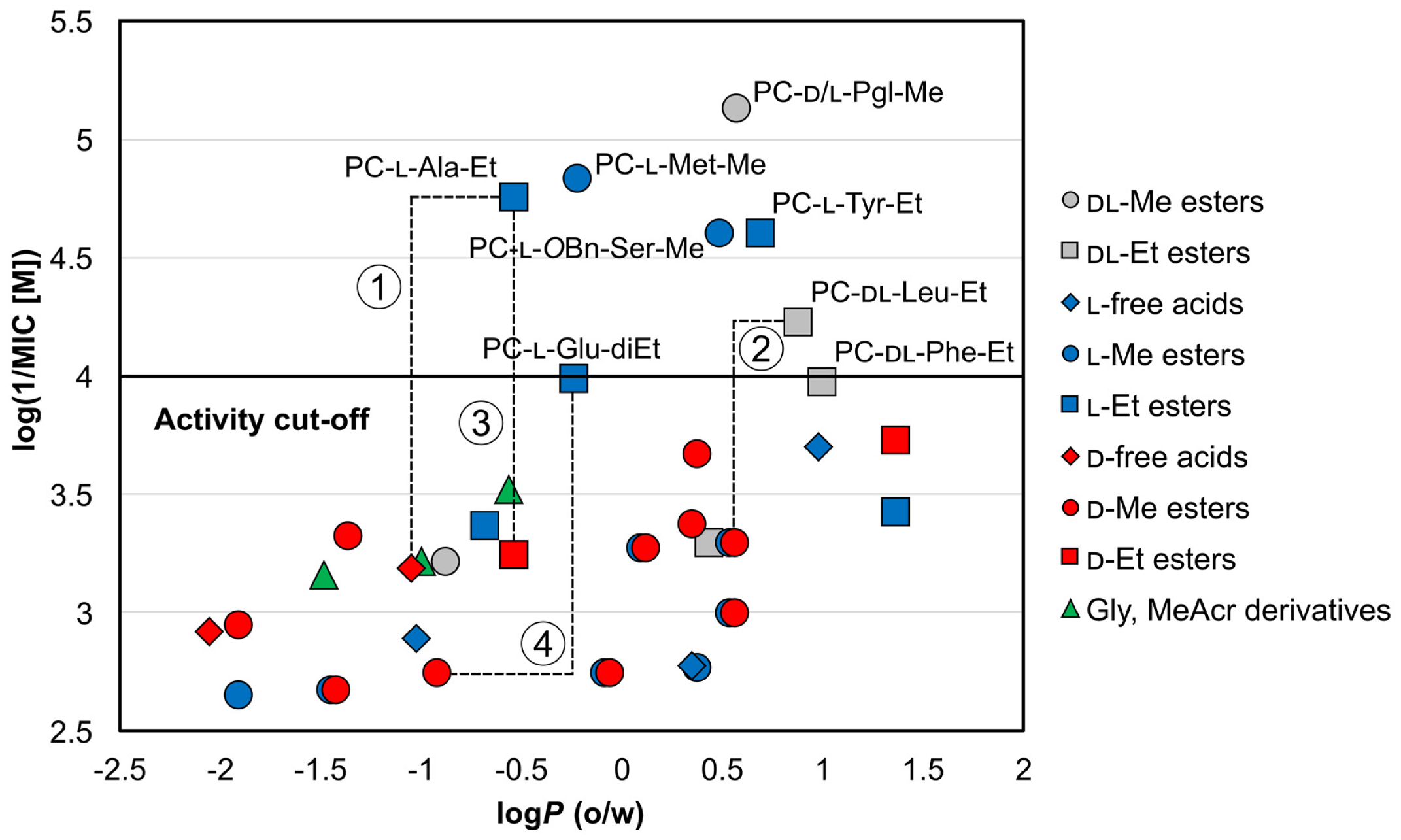
Molecules | Free Full-Text | N-Pyrazinoyl Substituted Amino Acids as Potential Antimycobacterial Agents—the Synthesis and Biological Evaluation of Enantiomers | HTML

Automated Parametrization of Biomolecular Force Fields from Quantum Mechanics/Molecular Mechanics (QM/MM) Simulations through Force Matching | Journal of Chemical Theory and Computation

Automated Parametrization of Biomolecular Force Fields from Quantum Mechanics/Molecular Mechanics (QM/MM) Simulations through Force Matching | Journal of Chemical Theory and Computation

X-ray screening identifies active site and allosteric inhibitors of SARS-CoV-2 main protease | Science

Pyrazinamide Susceptibility Is Driven by Activation of the SigE-Dependent Cell Envelope Stress Response in Mycobacterium tuberculosis | mBio